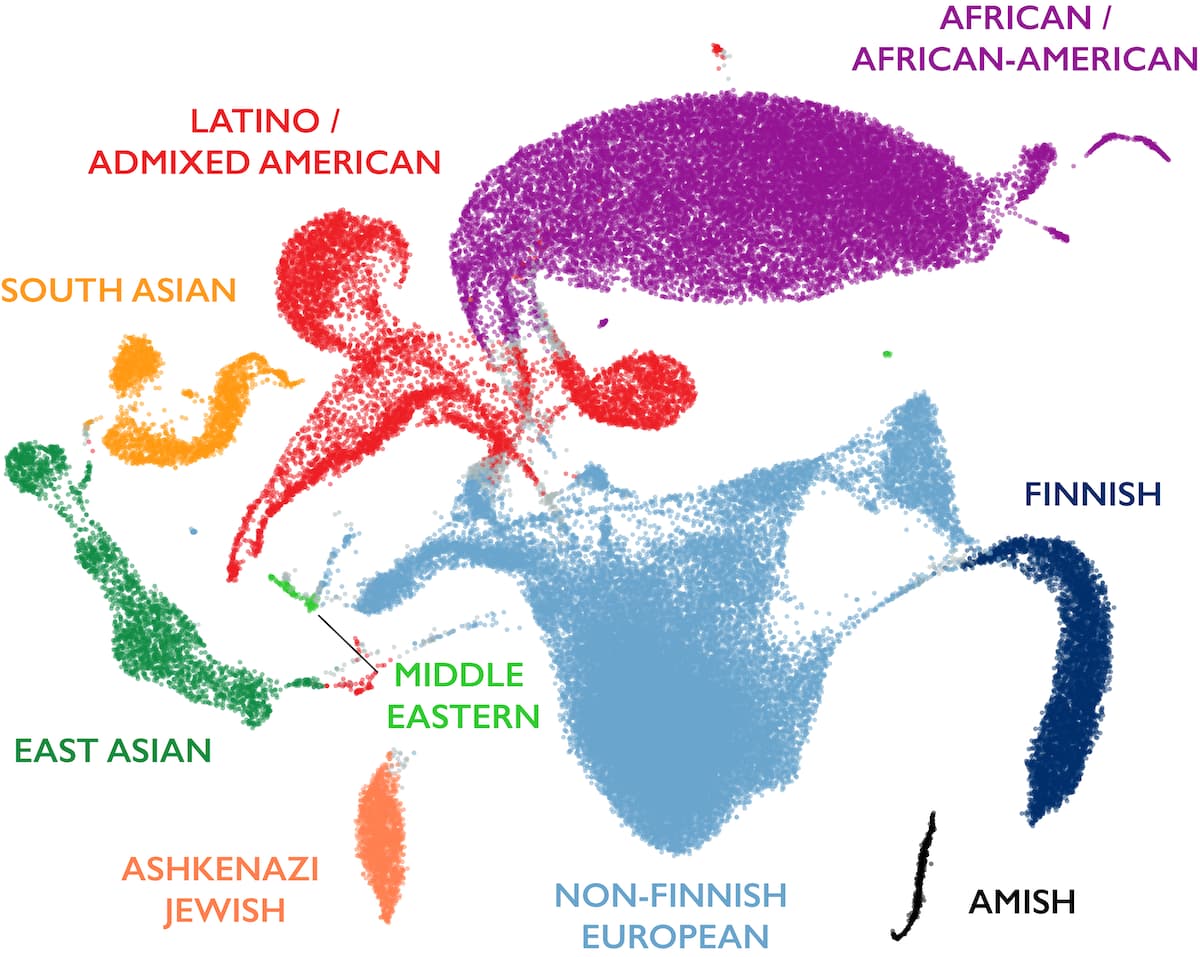
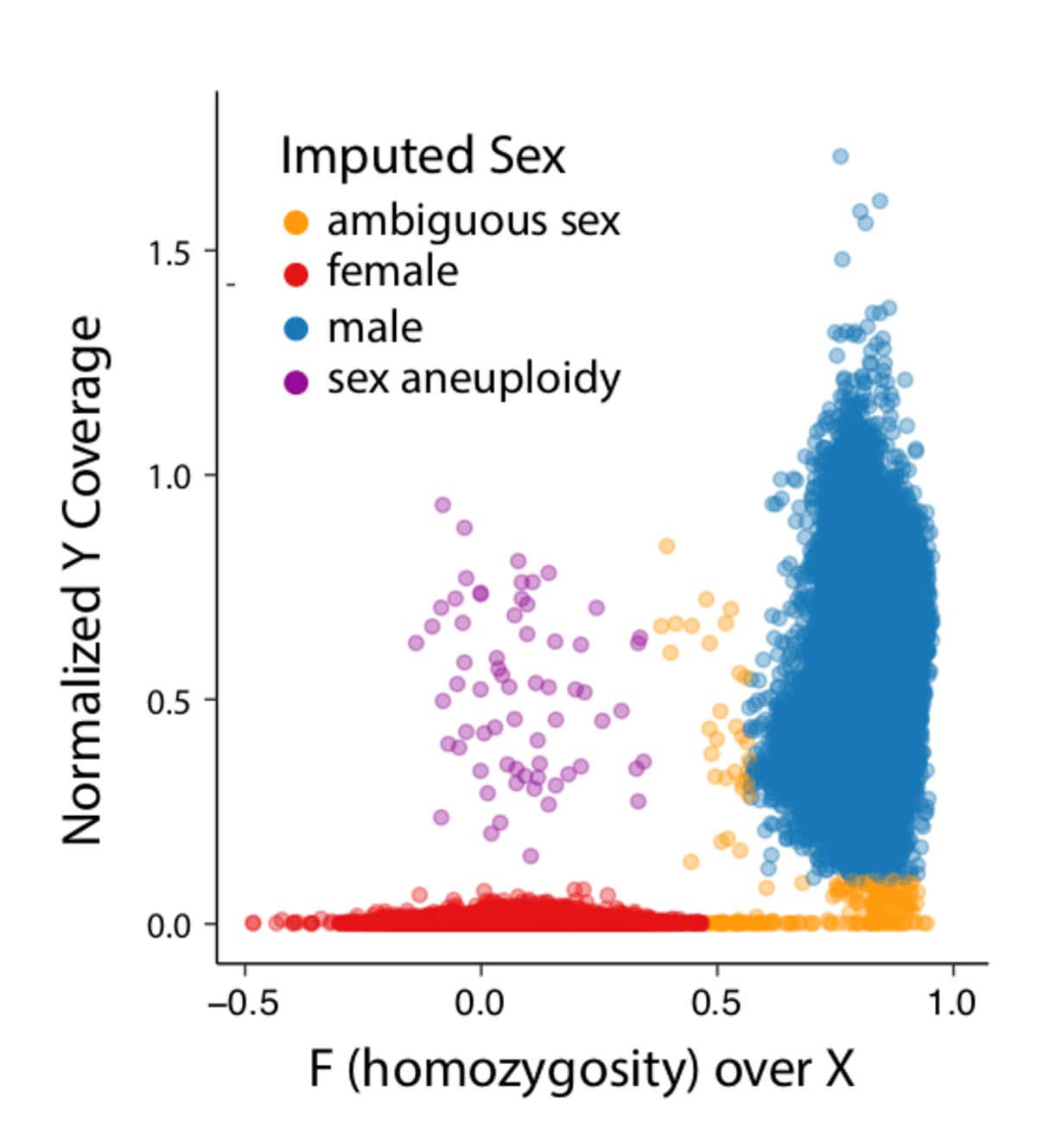
Variant interpretation using population databases: Lessons from gnomAD - Gudmundsson - 2022 - Human Mutation - Wiley Online Library

Systematic evaluation of gene variants linked to hearing loss based on allele frequency threshold and filtering allele frequency | Scientific Reports

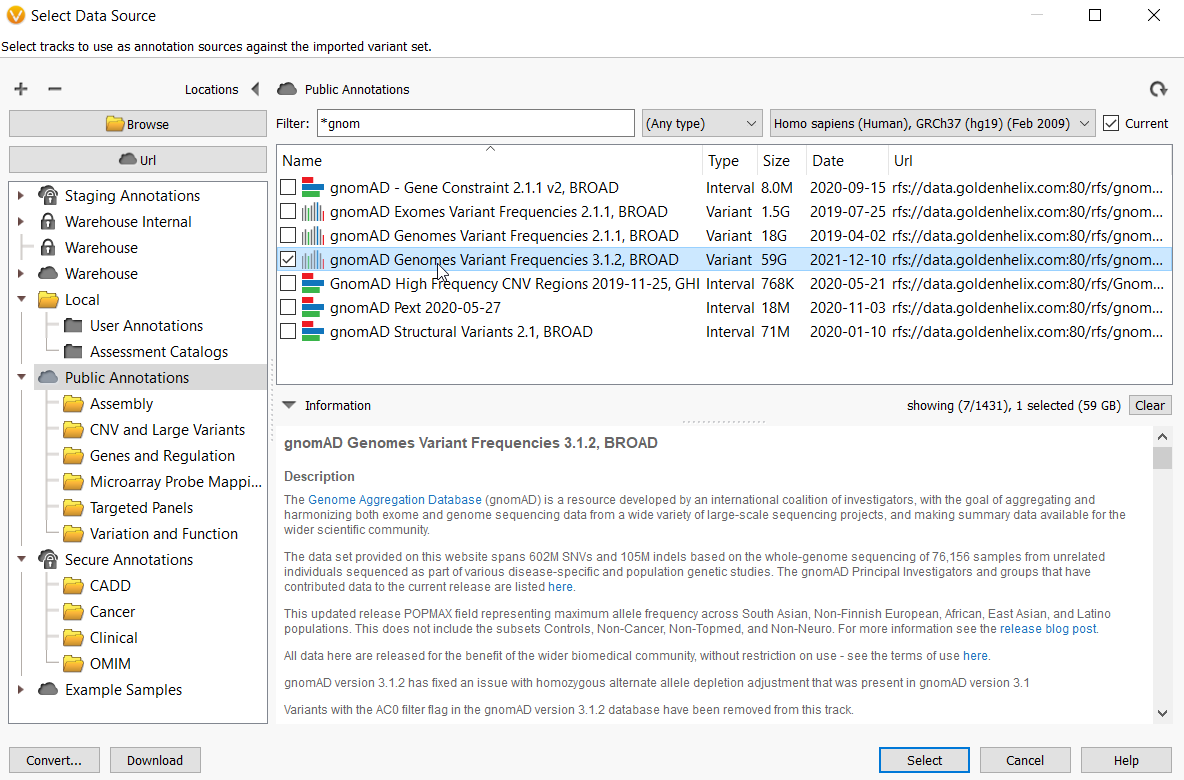
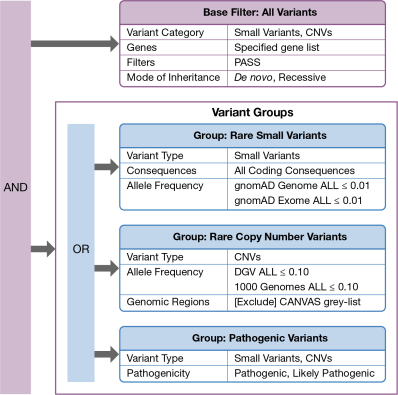
Using the Interactive Filter Cascade in QCI Interpret Translational - Bioinformatics Software | QIAGEN Digital Insights

Variant interpretation using population databases: Lessons from gnomAD - Gudmundsson - 2022 - Human Mutation - Wiley Online Library
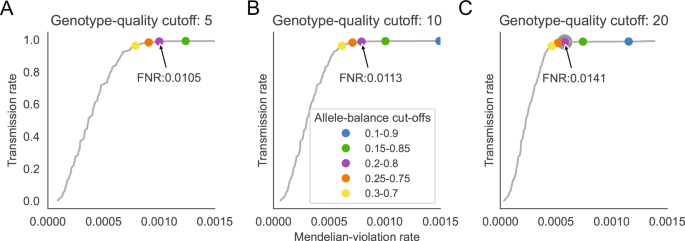
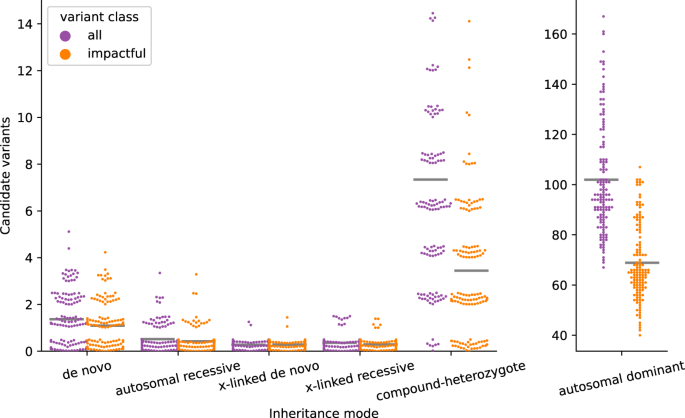
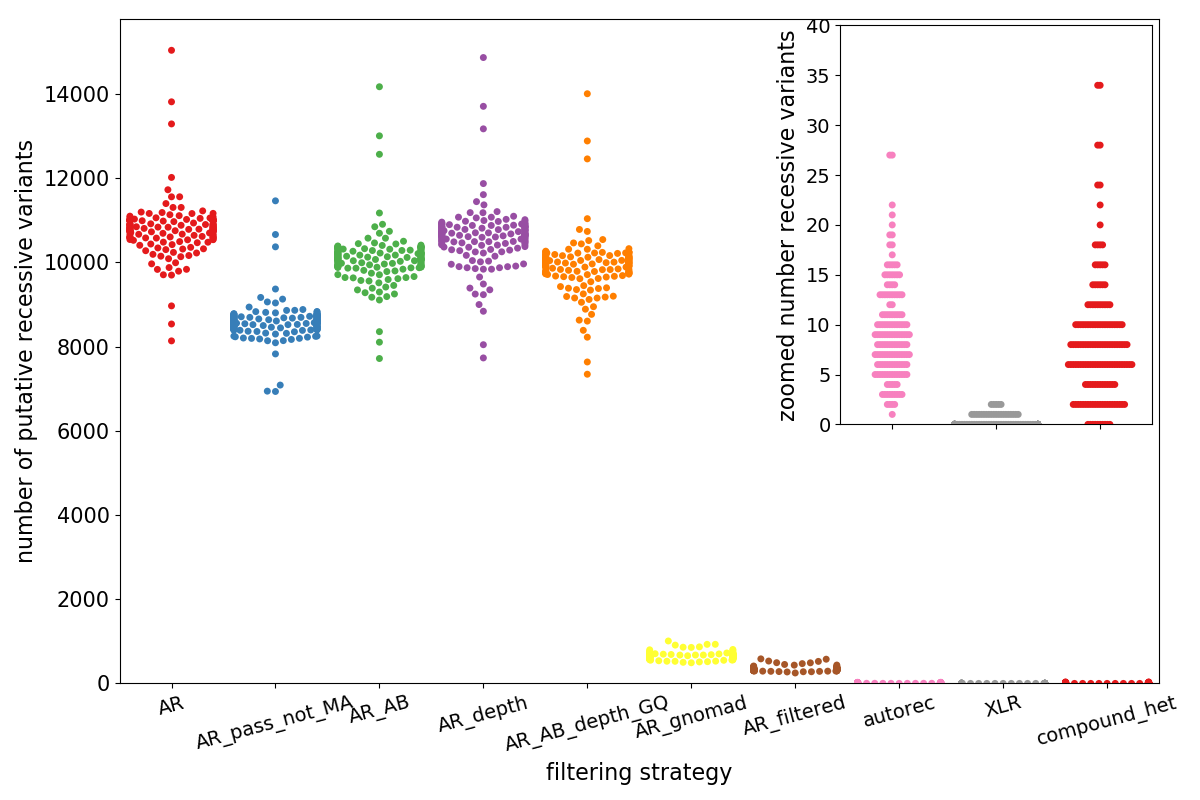
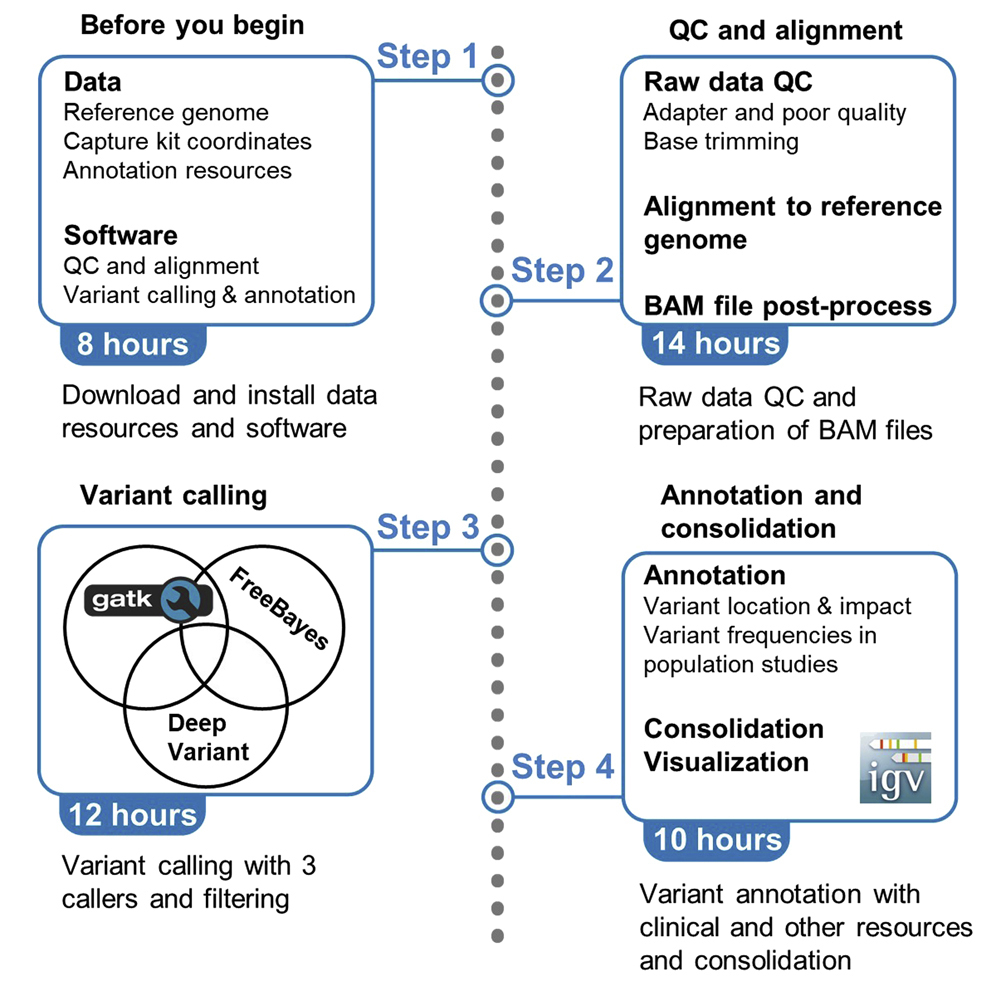
Effective variant filtering and expected candidate variant yield in studies of rare human disease | npj Genomic Medicine
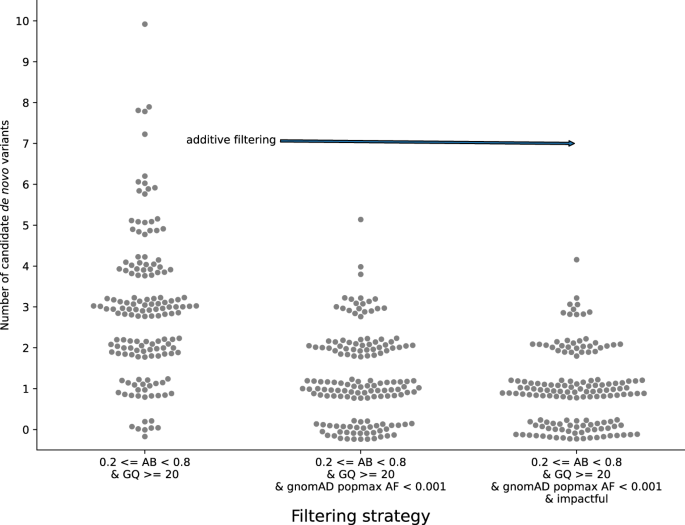
Effective variant filtering and expected candidate variant yield in studies of rare human disease | npj Genomic Medicine

Genome Aggregation Database on X: "We heard your feedback and updated the way #ClinVar data is displayed on #gnomAD browser. The ClinVar Track now includes: - Displaying variant type through shapes -
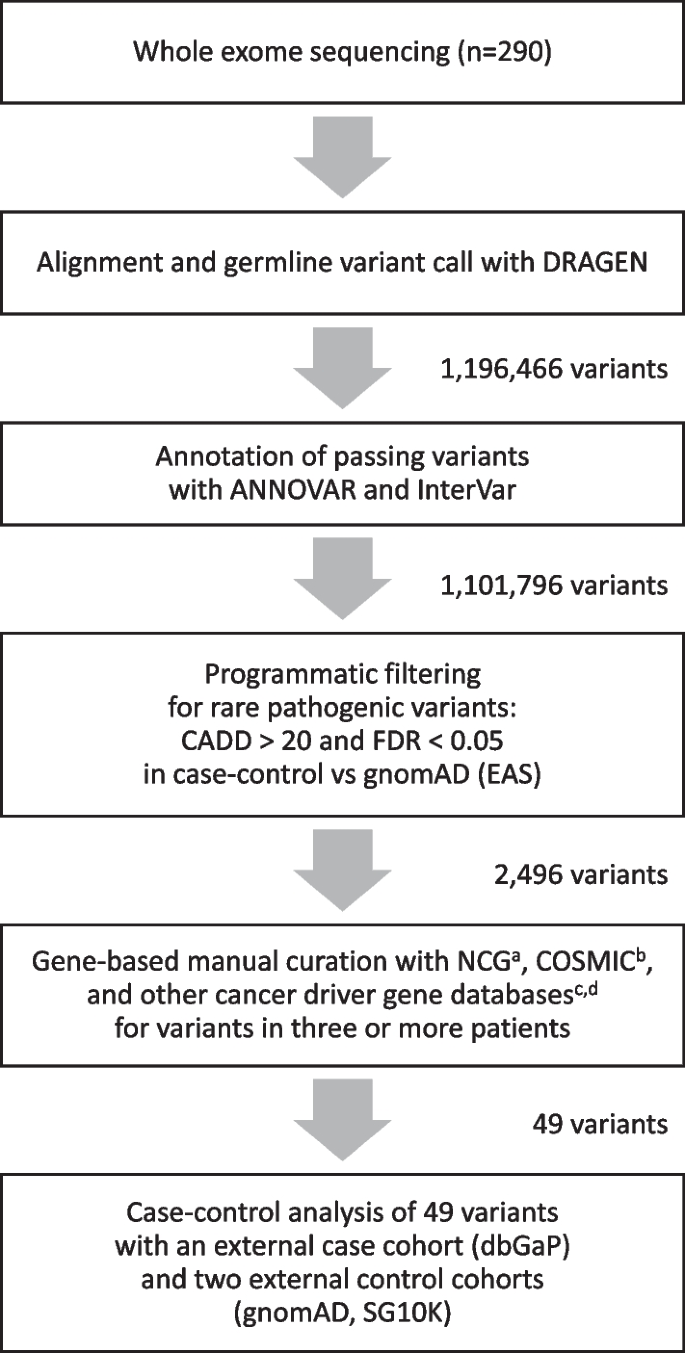
Whole-exome sequencing of BRCA-negative breast cancer patients and case–control analyses identify variants associated with breast cancer susceptibility | Human Genomics | Full Text
Comparison of observed and gnomAD filtering allele frequencies (AFs)... | Download Scientific Diagram

Genome Aggregation Database on X: "Another #gnomAD browser update! On variant pages you can now quickly switch between genome builds and gnomAD versions using our new liftover feature. This is available on

Adopting High-Resolution Allele Frequencies Substantially Expedites Variant Interpretation in Genetic Diagnostic Laboratories - ScienceDirect

Effective variant filtering and expected candidate variant yield in studies of rare human disease | bioRxiv